Note
Click here to download the full example code
Faces dataset decompositions¶
This example applies to olivetti_faces different unsupervised
matrix decomposition (dimension reduction) methods from the module
sklearn.decomposition (see the documentation chapter
Decomposing signals in components (matrix factorization problems)) .
Out:
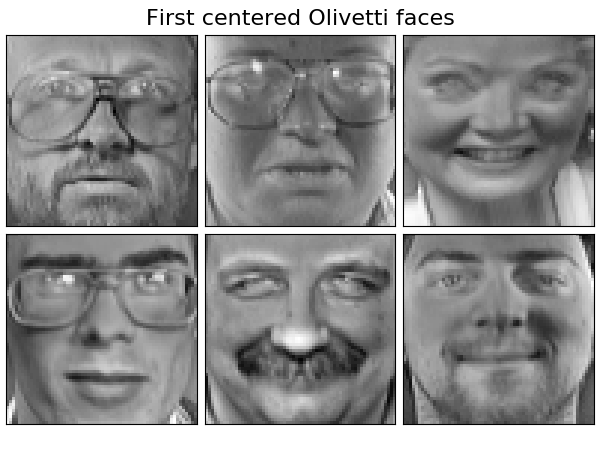
Dataset consists of 400 faces
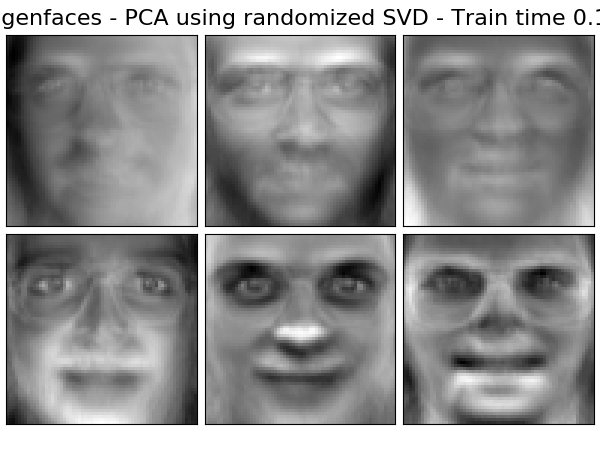
Extracting the top 6 Eigenfaces - PCA using randomized SVD...
done in 0.066s
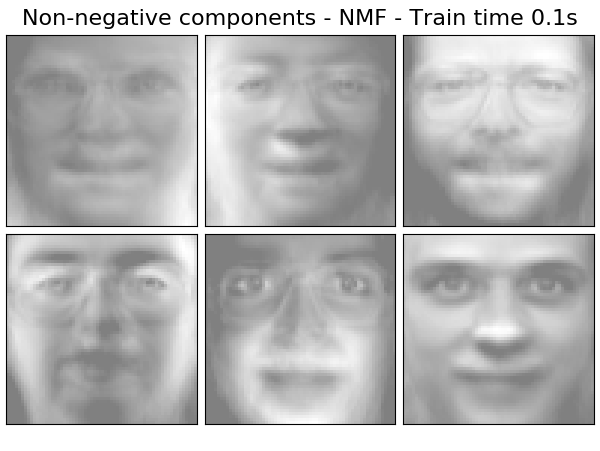
Extracting the top 6 Non-negative components - NMF...
done in 0.140s
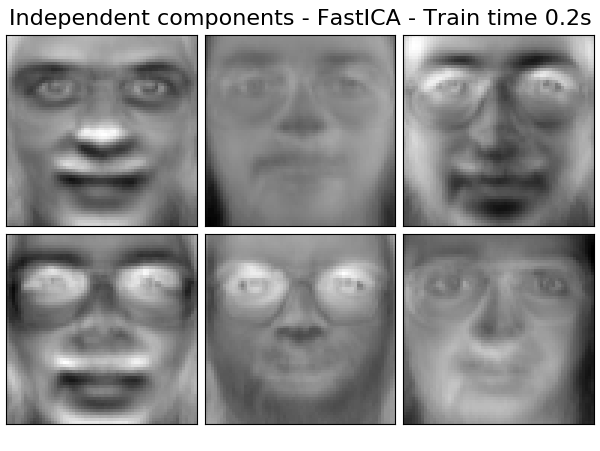
Extracting the top 6 Independent components - FastICA...
done in 0.206s
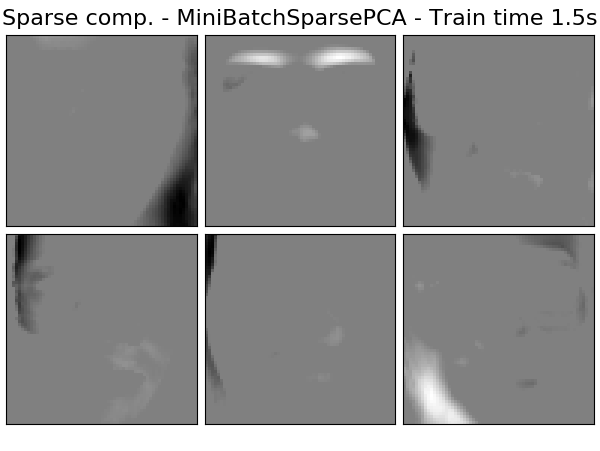
Extracting the top 6 Sparse comp. - MiniBatchSparsePCA...
done in 1.450s
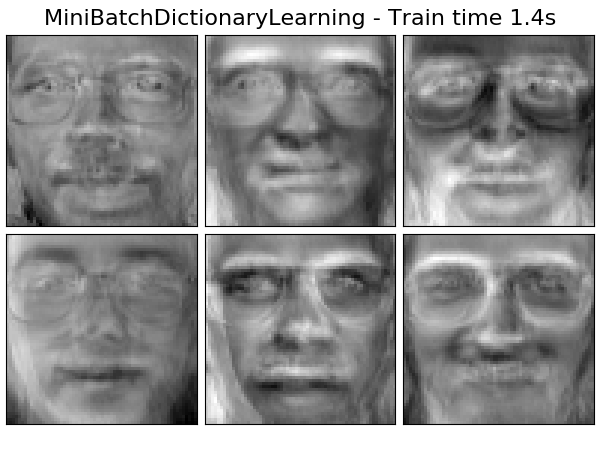
Extracting the top 6 MiniBatchDictionaryLearning...
done in 1.443s
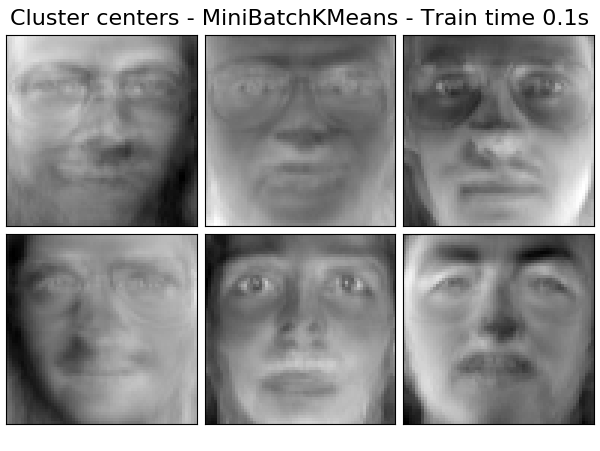
Extracting the top 6 Cluster centers - MiniBatchKMeans...
done in 0.145s
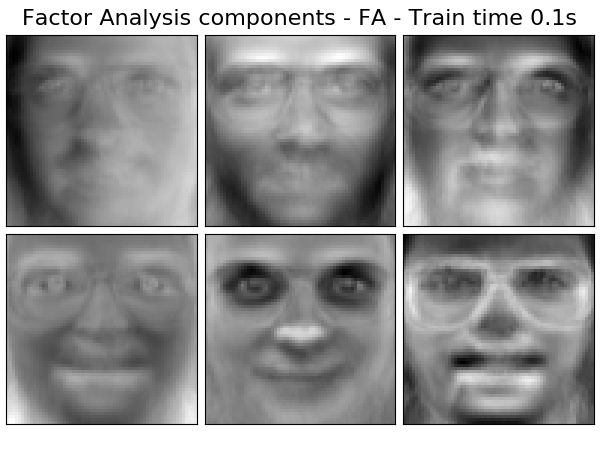
Extracting the top 6 Factor Analysis components - FA...
done in 0.105s
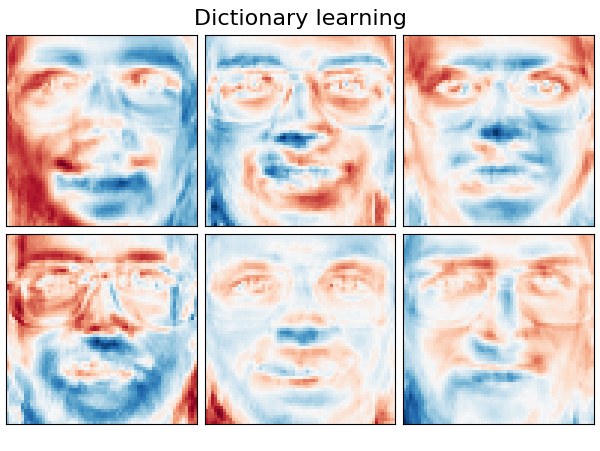
Extracting the top 6 Dictionary learning...
done in 1.553s
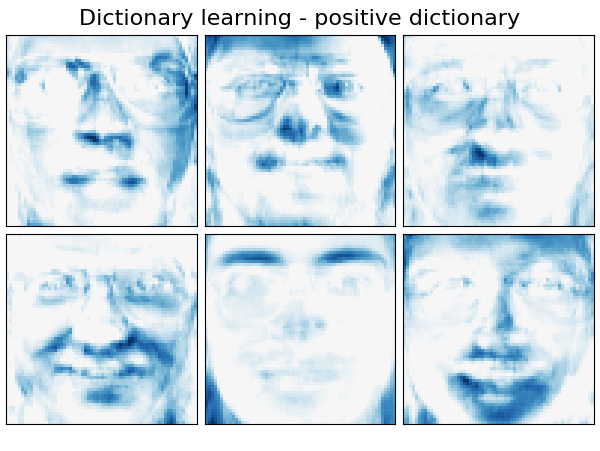
Extracting the top 6 Dictionary learning - positive dictionary...
done in 1.256s
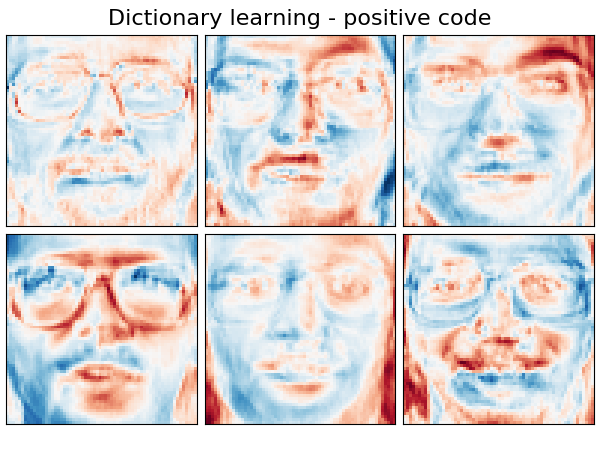
Extracting the top 6 Dictionary learning - positive code...
done in 0.734s
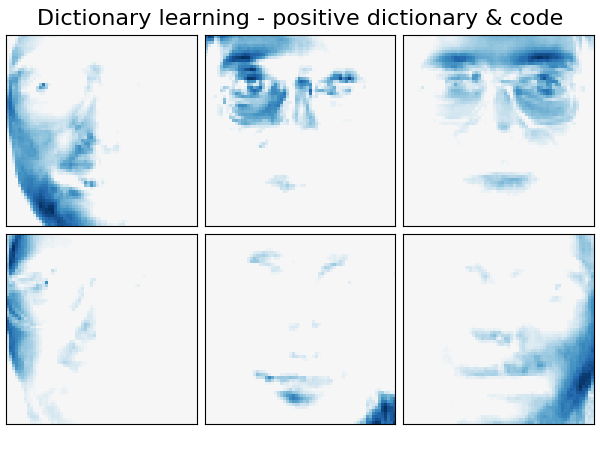
Extracting the top 6 Dictionary learning - positive dictionary & code...
done in 0.663s
print(__doc__)
# Authors: Vlad Niculae, Alexandre Gramfort
# License: BSD 3 clause
import logging
from time import time
from numpy.random import RandomState
import matplotlib.pyplot as plt
from sklearn.datasets import fetch_olivetti_faces
from sklearn.cluster import MiniBatchKMeans
from sklearn import decomposition
# Display progress logs on stdout
logging.basicConfig(level=logging.INFO,
format='%(asctime)s %(levelname)s %(message)s')
n_row, n_col = 2, 3
n_components = n_row * n_col
image_shape = (64, 64)
rng = RandomState(0)
# #############################################################################
# Load faces data
dataset = fetch_olivetti_faces(shuffle=True, random_state=rng)
faces = dataset.data
n_samples, n_features = faces.shape
# global centering
faces_centered = faces - faces.mean(axis=0)
# local centering
faces_centered -= faces_centered.mean(axis=1).reshape(n_samples, -1)
print("Dataset consists of %d faces" % n_samples)
def plot_gallery(title, images, n_col=n_col, n_row=n_row, cmap=plt.cm.gray):
plt.figure(figsize=(2. * n_col, 2.26 * n_row))
plt.suptitle(title, size=16)
for i, comp in enumerate(images):
plt.subplot(n_row, n_col, i + 1)
vmax = max(comp.max(), -comp.min())
plt.imshow(comp.reshape(image_shape), cmap=cmap,
interpolation='nearest',
vmin=-vmax, vmax=vmax)
plt.xticks(())
plt.yticks(())
plt.subplots_adjust(0.01, 0.05, 0.99, 0.93, 0.04, 0.)
# #############################################################################
# List of the different estimators, whether to center and transpose the
# problem, and whether the transformer uses the clustering API.
estimators = [
('Eigenfaces - PCA using randomized SVD',
decomposition.PCA(n_components=n_components, svd_solver='randomized',
whiten=True),
True),
('Non-negative components - NMF',
decomposition.NMF(n_components=n_components, init='nndsvda', tol=5e-3),
False),
('Independent components - FastICA',
decomposition.FastICA(n_components=n_components, whiten=True),
True),
('Sparse comp. - MiniBatchSparsePCA',
decomposition.MiniBatchSparsePCA(n_components=n_components, alpha=0.8,
n_iter=100, batch_size=3,
random_state=rng,
normalize_components=True),
True),
('MiniBatchDictionaryLearning',
decomposition.MiniBatchDictionaryLearning(n_components=15, alpha=0.1,
n_iter=50, batch_size=3,
random_state=rng),
True),
('Cluster centers - MiniBatchKMeans',
MiniBatchKMeans(n_clusters=n_components, tol=1e-3, batch_size=20,
max_iter=50, random_state=rng),
True),
('Factor Analysis components - FA',
decomposition.FactorAnalysis(n_components=n_components, max_iter=2),
True),
]
# #############################################################################
# Plot a sample of the input data
plot_gallery("First centered Olivetti faces", faces_centered[:n_components])
# #############################################################################
# Do the estimation and plot it
for name, estimator, center in estimators:
print("Extracting the top %d %s..." % (n_components, name))
t0 = time()
data = faces
if center:
data = faces_centered
estimator.fit(data)
train_time = (time() - t0)
print("done in %0.3fs" % train_time)
if hasattr(estimator, 'cluster_centers_'):
components_ = estimator.cluster_centers_
else:
components_ = estimator.components_
# Plot an image representing the pixelwise variance provided by the
# estimator e.g its noise_variance_ attribute. The Eigenfaces estimator,
# via the PCA decomposition, also provides a scalar noise_variance_
# (the mean of pixelwise variance) that cannot be displayed as an image
# so we skip it.
if (hasattr(estimator, 'noise_variance_') and
estimator.noise_variance_.ndim > 0): # Skip the Eigenfaces case
plot_gallery("Pixelwise variance",
estimator.noise_variance_.reshape(1, -1), n_col=1,
n_row=1)
plot_gallery('%s - Train time %.1fs' % (name, train_time),
components_[:n_components])
plt.show()
# #############################################################################
# Various positivity constraints applied to dictionary learning.
estimators = [
('Dictionary learning',
decomposition.MiniBatchDictionaryLearning(n_components=15, alpha=0.1,
n_iter=50, batch_size=3,
random_state=rng),
True),
('Dictionary learning - positive dictionary',
decomposition.MiniBatchDictionaryLearning(n_components=15, alpha=0.1,
n_iter=50, batch_size=3,
random_state=rng,
positive_dict=True),
True),
('Dictionary learning - positive code',
decomposition.MiniBatchDictionaryLearning(n_components=15, alpha=0.1,
n_iter=50, batch_size=3,
random_state=rng,
positive_code=True),
True),
('Dictionary learning - positive dictionary & code',
decomposition.MiniBatchDictionaryLearning(n_components=15, alpha=0.1,
n_iter=50, batch_size=3,
random_state=rng,
positive_dict=True,
positive_code=True),
True),
]
# #############################################################################
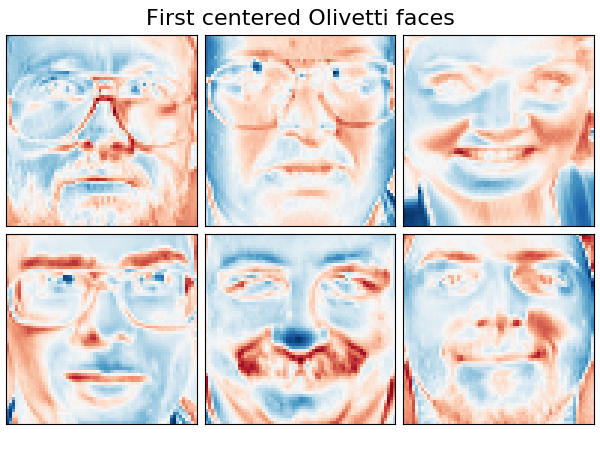
# Plot a sample of the input data
plot_gallery("First centered Olivetti faces", faces_centered[:n_components],
cmap=plt.cm.RdBu)
# #############################################################################
# Do the estimation and plot it
for name, estimator, center in estimators:
print("Extracting the top %d %s..." % (n_components, name))
t0 = time()
data = faces
if center:
data = faces_centered
estimator.fit(data)
train_time = (time() - t0)
print("done in %0.3fs" % train_time)
components_ = estimator.components_
plot_gallery(name, components_[:n_components], cmap=plt.cm.RdBu)
plt.show()
Total running time of the script: ( 0 minutes 9.759 seconds)