Non Linear Methods for Classification¶
Parzen-based classifier¶
- class mlpy.Parzen(kernel=None)¶
Parzen based classifier (binary).
Initialization.
Parameters : - kernel : None or mlpy.Kernel object.
if kernel is None, K and Kt in .learn() and in .pred() methods must be precomputed kernel matricies, else K and Kt must be training (resp. test) data in input space.
- alpha()¶
Return alpha.
- b()¶
Return b.
- labels()¶
Outputs the name of labels.
- learn(K, y)¶
Compute alpha and b.
- Parameters:
- K: 2d array_like object
- precomputed training kernel matrix (if kernel=None); training data in input space (if kernel is a Kernel object)
- y : 1d array_like object (N)
- target values
- pred(Kt)¶
Compute the predicted class.
Parameters : - Kt : 1d or 2d array_like object
precomputed test kernel matrix. (if kernel=None); test data in input space (if kernel is a Kernel object).
Returns : - p : integer or 1d numpy array
predicted class
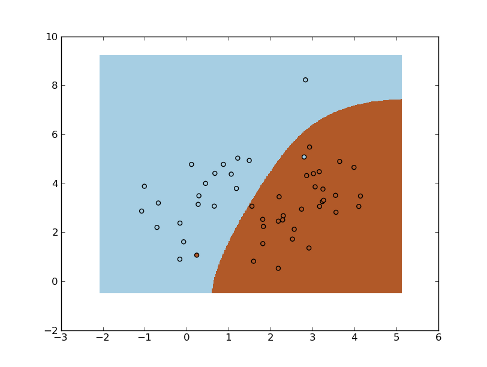
Example:
>>> import numpy as np
>>> import matplotlib.pyplot as plt
>>> import mlpy
>>> np.random.seed(0)
>>> mean1, cov1, n1 = [1, 4.5], [[1,1],[1,2]], 20 # 20 samples of class 1
>>> x1 = np.random.multivariate_normal(mean1, cov1, n1)
>>> y1 = np.ones(n1, dtype=np.int)
>>> mean2, cov2, n2 = [2.5, 2.5], [[1,1],[1,2]], 30 # 30 samples of class 2
>>> x2 = np.random.multivariate_normal(mean2, cov2, n2)
>>> y2 = 2 * np.ones(n2, dtype=np.int)
>>> x = np.concatenate((x1, x2), axis=0) # concatenate the samples
>>> y = np.concatenate((y1, y2))
>>> K = mlpy.kernel_gaussian(x, x, sigma=2) # kernel matrix
>>> parzen = mlpy.Parzen()
>>> parzen.learn(K, y)
>>> xmin, xmax = x[:,0].min()-1, x[:,0].max()+1
>>> ymin, ymax = x[:,1].min()-1, x[:,1].max()+1
>>> xx, yy = np.meshgrid(np.arange(xmin, xmax, 0.02), np.arange(ymin, ymax, 0.02))
>>> xt = np.c_[xx.ravel(), yy.ravel()] # test points
>>> Kt = mlpy.kernel_gaussian(xt, x, sigma=2) # test kernel matrix
>>> yt = parzen.pred(Kt).reshape(xx.shape)
>>> fig = plt.figure(1)
>>> cmap = plt.set_cmap(plt.cm.Paired)
>>> plot1 = plt.pcolormesh(xx, yy, yt)
>>> plot2 = plt.scatter(x[:,0], x[:,1], c=y)
>>> plt.show()
Support Vector Classification¶
Kernel Fisher Discriminant Classifier¶
- class mlpy.KFDAC(lmb=0.001, kernel=None)¶
Kernel Fisher Discriminant Analysis Classifier (binary classifier).
The bias term (b) is computed as in [Gavin03].
[Gavin03] Gavin C. et al. Efficient Cross-Validation of Kernel Fisher Discriminant Classifers. ESANN‘2003 proceedings - European Symposium on Artificial Neural Networks, 2003. Initialization.
Parameters : - lmb : float (>= 0.0)
regularization parameter
- kernel : None or mlpy.Kernel object.
if kernel is None, K and Kt in .learn() and in .transform() methods must be precomputed kernel matricies, else K and Kt must be training (resp. test) data in input space.
- alpha()¶
Return alpha.
- b()¶
Return b.
- labels()¶
Outputs the name of labels.
- learn(K, y)¶
Learning method.
Parameters : - K: 2d array_like object
precomputed training kernel matrix (if kernel=None); training data in input space (if kernel is a Kernel object)
- y : 1d array_like object integer (N)
class labels (only two classes)
- pred(Kt)¶
Compute the predicted response.
Parameters : - Kt : 1d or 2d array_like object
precomputed test kernel matrix. (if kernel=None); test data in input space (if kernel is a Kernel object).
Returns : - p : integer or 1d numpy array
the predicted class(es)
k-Nearest-Neighbor¶
- class mlpy.KNN(k)¶
k-Nearest Neighbor (euclidean distance)
Parameters : - k : int
number of nearest neighbors
- KNN.learn(x, y)¶
Learn method.
Parameters : - x : 2d array_like object (N,P)
training data
- y : 1d array_like integer
class labels
- KNN.pred(t)¶
Predict KNN model on a test point(s).
Parameters : - t : 1d or 2d array_like object ([M,] P)
test point(s)
Returns : - p : int or 1d numpy array
the predicted value(s). Retuns the smallest label minus one (KNN.labels()[0]-1) when the classification is not unique.
- KNN.nclasses()¶
Returns the number of classes.
- KNN.labels()¶
Outputs the name of labels.
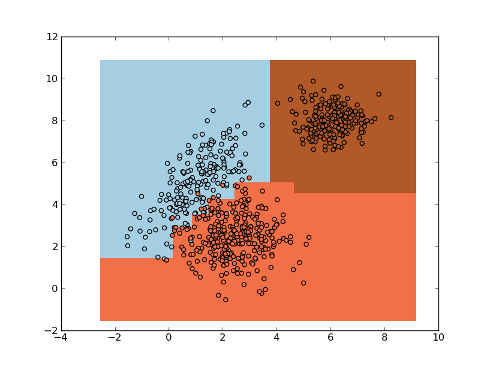
Example:
>>> import numpy as np
>>> import matplotlib.pyplot as plt
>>> import mlpy
>>> np.random.seed(0)
>>> mean1, cov1, n1 = [1, 5], [[1,1],[1,2]], 200 # 200 samples of class 1
>>> x1 = np.random.multivariate_normal(mean1, cov1, n1)
>>> y1 = np.ones(n1, dtype=np.int)
>>> mean2, cov2, n2 = [2.5, 2.5], [[1,0],[0,1]], 300 # 300 samples of class 2
>>> x2 = np.random.multivariate_normal(mean2, cov2, n2)
>>> y2 = 2 * np.ones(n2, dtype=np.int)
>>> mean3, cov3, n3 = [5, 8], [[0.5,0],[0,0.5]], 200 # 200 samples of class 3
>>> x3 = np.random.multivariate_normal(mean3, cov3, n3)
>>> y3 = 3 * np.ones(n3, dtype=np.int)
>>> x = np.concatenate((x1, x2, x3), axis=0) # concatenate the samples
>>> y = np.concatenate((y1, y2, y3))
>>> knn = mlpy.KNN(k=3)
>>> knn.learn(x, y)
>>> xmin, xmax = x[:,0].min()-1, x[:,0].max()+1
>>> ymin, ymax = x[:,1].min()-1, x[:,1].max()+1
>>> xx, yy = np.meshgrid(np.arange(xmin, xmax, 0.1), np.arange(ymin, ymax, 0.1))
>>> xnew = np.c_[xx.ravel(), yy.ravel()]
>>> ynew = knn.pred(xnew).reshape(xx.shape)
>>> ynew[ynew == 0] = 1 # set the samples with no unique classification to 1
>>> fig = plt.figure(1)
>>> cmap = plt.set_cmap(plt.cm.Paired)
>>> plot1 = plt.pcolormesh(xx, yy, ynew)
>>> plot2 = plt.scatter(x[:,0], x[:,1], c=y)
>>> plt.show()
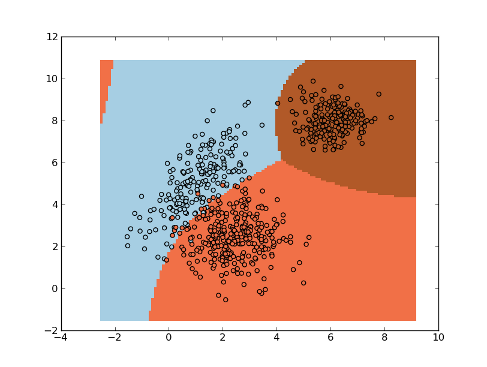
Classification Tree¶
- class mlpy.ClassTree(stumps=0, minsize=1)¶
Classification Tree (gini index)
Parameters : - stumps : bool
True: compute single split or False: standard tree
- minsize : int (>=0)
minimum number of cases required to split a leaf
- ClassTree.learn(x, y)¶
Learn method.
Parameters : - x : 2d array_like object (N x P)
training data
- y : 1d array_like integer
class labels
- ClassTree.pred(t)¶
Predict Tree model on a test point(s).
Parameters : - t : 1d or 2d array_like object ([M,] P)
test point(s)
Returns : - p : int or 1d numpy array
the predicted value(s). Retuns the smallest label minus one (ClassTree.labels()[0]-1) when the classification is not unique.
- ClassTree.nclasses()¶
Returns the number of classes.
- ClassTree.labels()¶
Outputs the name of labels.
Example:
>>> import numpy as np
>>> import matplotlib.pyplot as plt
>>> import mlpy
>>> np.random.seed(0)
>>> mean1, cov1, n1 = [1, 5], [[1,1],[1,2]], 200 # 200 samples of class 1
>>> x1 = np.random.multivariate_normal(mean1, cov1, n1)
>>> y1 = np.ones(n1, dtype=np.int)
>>> mean2, cov2, n2 = [2.5, 2.5], [[1,0],[0,1]], 300 # 300 samples of class 2
>>> x2 = np.random.multivariate_normal(mean2, cov2, n2)
>>> y2 = 2 * np.ones(n2, dtype=np.int)
>>> mean3, cov3, n3 = [6, 8], [[0.5,0],[0,0.5]], 200 # 200 samples of class 3
>>> x3 = np.random.multivariate_normal(mean3, cov3, n3)
>>> y3 = 3 * np.ones(n3, dtype=np.int)
>>> x = np.concatenate((x1, x2, x3), axis=0) # concatenate the samples
>>> y = np.concatenate((y1, y2, y3))
>>> tree = mlpy.ClassTree(minsize=10)
>>> tree.learn(x, y)
>>> xmin, xmax = x[:,0].min()-1, x[:,0].max()+1
>>> ymin, ymax = x[:,1].min()-1, x[:,1].max()+1
>>> xx, yy = np.meshgrid(np.arange(xmin, xmax, 0.1), np.arange(ymin, ymax, 0.1))
>>> xnew = np.c_[xx.ravel(), yy.ravel()]
>>> ynew = tree.pred(xnew).reshape(xx.shape)
>>> ynew[ynew == 0] = 1 # set the samples with no unique classification to 1
>>> fig = plt.figure(1)
>>> cmap = plt.set_cmap(plt.cm.Paired)
>>> plot1 = plt.pcolormesh(xx, yy, ynew)
>>> plot2 = plt.scatter(x[:,0], x[:,1], c=y)
>>> plt.show()
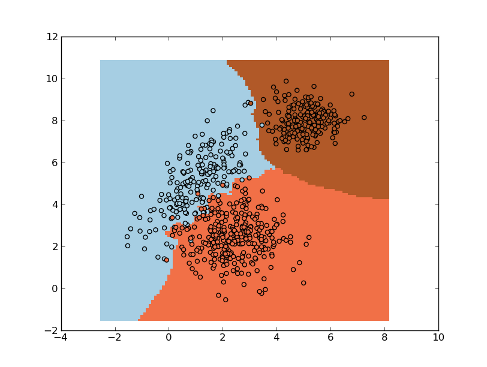
Maximum Likelihood Classifier¶
- class mlpy.MaximumLikelihoodC¶
Maximum Likelihood Classifier
- MaximumLikelihoodC.learn(x, y)¶
Learn method.
Parameters : - x : 2d array_like object (N,P)
training data
- y : 1d array_like integer
class labels
- MaximumLikelihoodC.pred(t)¶
Predict Maximum Likelihood model on a test point(s).
Parameters : - t : 1d or 2d array_like object ([M,] P)
test point(s)
Returns : - p : int or 1d numpy array
the predicted value(s). Retuns the smallest label minus one (MaximumLikelihoodC.labels()[0]-1) when the classification is not unique.
- MaximumLikelihoodC.nclasses()¶
Returns the number of classes.
- MaximumLikelihoodC.labels()¶
Outputs the name of labels.
Example:
>>> import numpy as np
>>> import matplotlib.pyplot as plt
>>> import mlpy
>>> np.random.seed(0)
>>> mean1, cov1, n1 = [1, 5], [[1,1],[1,2]], 200 # 200 samples of class 1
>>> x1 = np.random.multivariate_normal(mean1, cov1, n1)
>>> y1 = np.ones(n1, dtype=np.int)
>>> mean2, cov2, n2 = [2.5, 2.5], [[1,0],[0,1]], 300 # 300 samples of class 2
>>> x2 = np.random.multivariate_normal(mean2, cov2, n2)
>>> y2 = 2 * np.ones(n2, dtype=np.int)
>>> mean3, cov3, n3 = [6, 8], [[0.5,0],[0,0.5]], 200 # 200 samples of class 3
>>> x3 = np.random.multivariate_normal(mean3, cov3, n3)
>>> y3 = 3 * np.ones(n3, dtype=np.int)
>>> x = np.concatenate((x1, x2, x3), axis=0) # concatenate the samples
>>> y = np.concatenate((y1, y2, y3))
>>> ml = mlpy.MaximumLikelihoodC()
>>> ml.learn(x, y)
>>> xmin, xmax = x[:,0].min()-1, x[:,0].max()+1
>>> ymin, ymax = x[:,1].min()-1, x[:,1].max()+1
>>> xx, yy = np.meshgrid(np.arange(xmin, xmax, 0.1), np.arange(ymin, ymax, 0.1))
>>> xnew = np.c_[xx.ravel(), yy.ravel()]
>>> ynew = ml.pred(xnew).reshape(xx.shape)
>>> ynew[ynew == 0] = 1 # set the samples with no unique classification to 1
>>> fig = plt.figure(1)
>>> cmap = plt.set_cmap(plt.cm.Paired)
>>> plot1 = plt.pcolormesh(xx, yy, ynew)
>>> plot2 = plt.scatter(x[:,0], x[:,1], c=y)
>>> plt.show()